|
probeID |
AGICode |
Annotation |
Log2 signal ratio
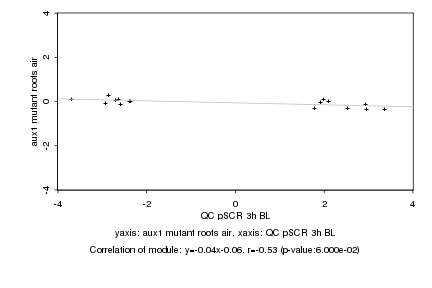
QC pSCR 3h BL |
Log2 signal ratio
aux1 mutant roots air |
| 1 |
AT1G52690 |
AT1G52690
|
LEA7, LATE EMBRYOGENESIS ABUNDANT 7 |
3.347 |
-0.346 |
| 2 |
AT4G12410 |
AT4G12410
|
SAUR35, SMALL AUXIN UPREGULATED RNA 35 |
2.934 |
-0.343 |
| 3 |
AT2G26150 |
AT2G26150
|
ATHSFA2, heat shock transcription factor A2, HSFA2, heat shock transcription factor A2 |
2.917 |
-0.113 |
| 4 |
AT1G29500 |
AT1G29500
|
SAUR66, SMALL AUXIN UPREGULATED RNA 66 |
2.521 |
-0.307 |
| 5 |
AT4G21745 |
AT4G21745
|
[PAK-box/P21-Rho-binding family protein] |
2.487 |
no data |
| 6 |
AT3G05936 |
AT3G05936
|
unknown |
2.303 |
no data |
| 7 |
AT2G31540 |
AT2G31540
|
[GDSL-like Lipase/Acylhydrolase superfamily protein] |
2.303 |
no data |
| 8 |
AT1G19530 |
AT1G19530
|
unknown |
2.088 |
0.034 |
| 9 |
AT1G26210 |
AT1G26210
|
ATSOFL1, SOB FIVE-LIKE 1, SOFL1, SOB five-like 1 |
1.978 |
0.117 |
| 10 |
AT4G21870 |
AT4G21870
|
[HSP20-like chaperones superfamily protein] |
1.916 |
-0.027 |
| 11 |
AT3G14370 |
AT3G14370
|
WAG2 |
1.767 |
-0.292 |
| 12 |
AT5G66080 |
AT5G66080
|
APD9, Arabidopsis Pp2c clade D 9 |
-3.711 |
0.119 |
| 13 |
AT1G68825 |
AT1G68825
|
DVL5, DEVIL 5, RTFL15, ROTUNDIFOLIA like 15 |
-3.125 |
no data |
| 14 |
AT1G28360 |
AT1G28360
|
ATERF12, ERF DOMAIN PROTEIN 12, ERF12, ERF domain protein 12 |
-2.935 |
-0.082 |
| 15 |
AT3G12900 |
AT3G12900
|
[2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein] |
-2.880 |
0.316 |
| 16 |
AT1G03800 |
AT1G03800
|
ATERF10, ARABIDOPSIS THALIANA RF DOMAIN PROTEIN 10, ERF10, ERF domain protein 10 |
-2.745 |
no data |
| 17 |
AT3G49630 |
AT3G49630
|
[2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein] |
-2.716 |
0.057 |
| 18 |
AT3G62760 |
AT3G62760
|
ATGSTF13 |
-2.650 |
0.101 |
| 19 |
AT5G22580 |
AT5G22580
|
[Stress responsive A/B Barrel Domain] |
-2.597 |
-0.111 |
| 20 |
AT1G26870 |
AT1G26870
|
ANAC009, Arabidopsis NAC domain containing protein 9, FEZ, FEZ |
-2.391 |
0.046 |
| 21 |
AT3G26120 |
AT3G26120
|
terminal EAR1-like 1 |
-2.377 |
0.005 |