| ID | C0053 |
| Compound name | ADP |
| Stereochemistry | - |
| Aracyc name | ADP |
| External link |
|
| Mass information |
1st experiment : 2nd experiment : CE-TOF/MS (Anion analysis) |
| Molecular Formula | H12P2N5C10O10 |
| Canonical SMILES | C3(N(C1(C(C(C(O1)COP(OP(=O)([O-])[O-])([O-])=O)O)O))C2(=C(C(=NC=N2)N)N=3)) |
| Molecular Weight | 427.203 |
| Pathway Information | phosphatidylcholine biosynthesis I, mannitol degradation II, lipid IVA biosynthesis, stachyose degradation, Rubisco shunt, copper transport II, glycerol degradation IV, chorismate biosynthesis, folate polyglutamylation II, galactose degradation III, D-myo-inositol (1,4,5)-trisphosphate biosynthesis, UDP-L-arabinose biosynthesis II (from L-arabinose), cadmium transport I, folate polyglutamylation, coenzyme A biosynthesis, sulfate activation for sulfonation, 1,4-dihydroxy-2-naphthoate biosynthesis II (plants), pyrimidine deoxyribonucleotides de novo biosynthesis I, galactose degradation II, inosine-5'-phosphate biosynthesis II, ribose degradation, acetyl-CoA biosynthesis (from citrate), inositol pyrophosphates biosynthesis, fatty acid biosynthesis initiation I, xylose degradation I, adenine and adenosine salvage VI, TCA cycle variation III (eukaryotic), tetrahydrofolate biosynthesis II, phosphatidylethanolamine biosynthesis II, pyridoxal 5'-phosphate salvage pathway, citrulline biosynthesis, trans-zeatin biosynthesis, glutamine biosynthesis I, arginine biosynthesis II (acetyl cycle), L-Ndelta-acetylornithine biosynthesis, mevalonate pathway I, methionine biosynthesis II, trehalose degradation II (trehalase), citrulline degradation, methionine salvage pathway, TCA cycle variation V (plant), chlorophyllide a biosynthesis I, starch degradation I, methylerythritol phosphate pathway, pyridine nucleotide cycling (plants), ceramide degradation, biotin-carboxyl carrier protein assembly, purine nucleotide metabolism (phosphotransfer and nucleotide modification), UDP-D-galacturonate biosynthesis II (from D-galacturonate), asparagine biosynthesis III (tRNA-dependent), superpathway of glyoxylate cycle and fatty acid degradation, NAD/NADH phosphorylation and dephosphorylation, 1D-myo-inositol hexakisphosphate biosynthesis V (from Ins(1,3,4)P3), galactose degradation I (Leloir pathway), uridine-5'-phosphate biosynthesis, lipid-dependent phytate biosynthesis II (via Ins(1,3,4)P3), sucrose degradation III, lysine biosynthesis VI, sphingolipid biosynthesis (plants), choline biosynthesis I, GDP-glucose biosynthesis, photorespiration, gamma-glutamyl cycle (plant pathway), 5-aminoimidazole ribonucleotide biosynthesis I, gamma-glutamyl cycle, thiamine biosynthesis II, proline biosynthesis III, glutathione biosynthesis, folate transformations II, ornithine biosynthesis, diphthamide biosynthesis, S-methyl-5'-thioadenosine degradation I, starch biosynthesis, arginine biosynthesis I, homoserine biosynthesis, flavin biosynthesis I (bacteria and plants), sucrose biosynthesis I, leucine degradation I, glycolysis I, pyrimidine ribonucleotides interconversion, 1D-myo-inositol hexakisphosphate biosynthesis III (Spirodela polyrrhiza), glycolysis IV (plant cytosol), Calvin-Benson-Bassham cycle, UDP-D-glucuronate biosynthesis (from myo-inositol), guanine and guanosine salvage III, glutamine biosynthesis III, L-glutamine biosynthesis II (tRNA-dependent), GDP-L-fucose biosynthesis II (from L-fucose), nitrate reduction II (assimilatory), ammonia assimilation cycle II, gluconeogenesis I, threonine biosynthesis from homoserine, mannose degradation, urea cycle, lipid-dependent phytate biosynthesis I (via Ins(1,4,5)P3), biotin biosynthesis II |
| CAS ID | 58-64-0, 20398-34-9 |
| Synonym | adenosine pyrophosphate; adenosine 5'-pyrophosphate; adenosine-5'-diphosphate; adenosine-diphosphate; adenosine-5-diphosphate |
| External ID | CHEBI:16761, CHEBI:456216, PUBCHEM:7058055, PUBCHEM:6022, PUBCHEM:7058055, LIGAND-CPD:C00008 |


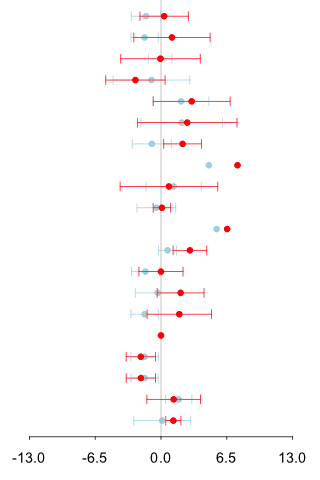

information